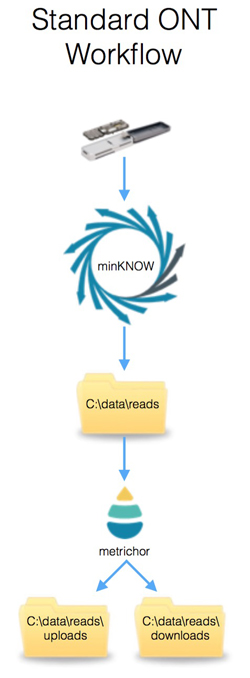
Jan Gawor on Twitter: "After 29h of sequencing: MinKnow 2 prediction: 13GB My calculation from initial data QC -> 11GB😃 After remux still 300 pores in strand 😁😁😁" / Twitter

Oxford Nanopore on Twitter: "SR: a reminder that we have the MinKNOW app, to support #anythinganyoneanywhere #nanoporeconf https://t.co/C6qXNkAkT3" / Twitter
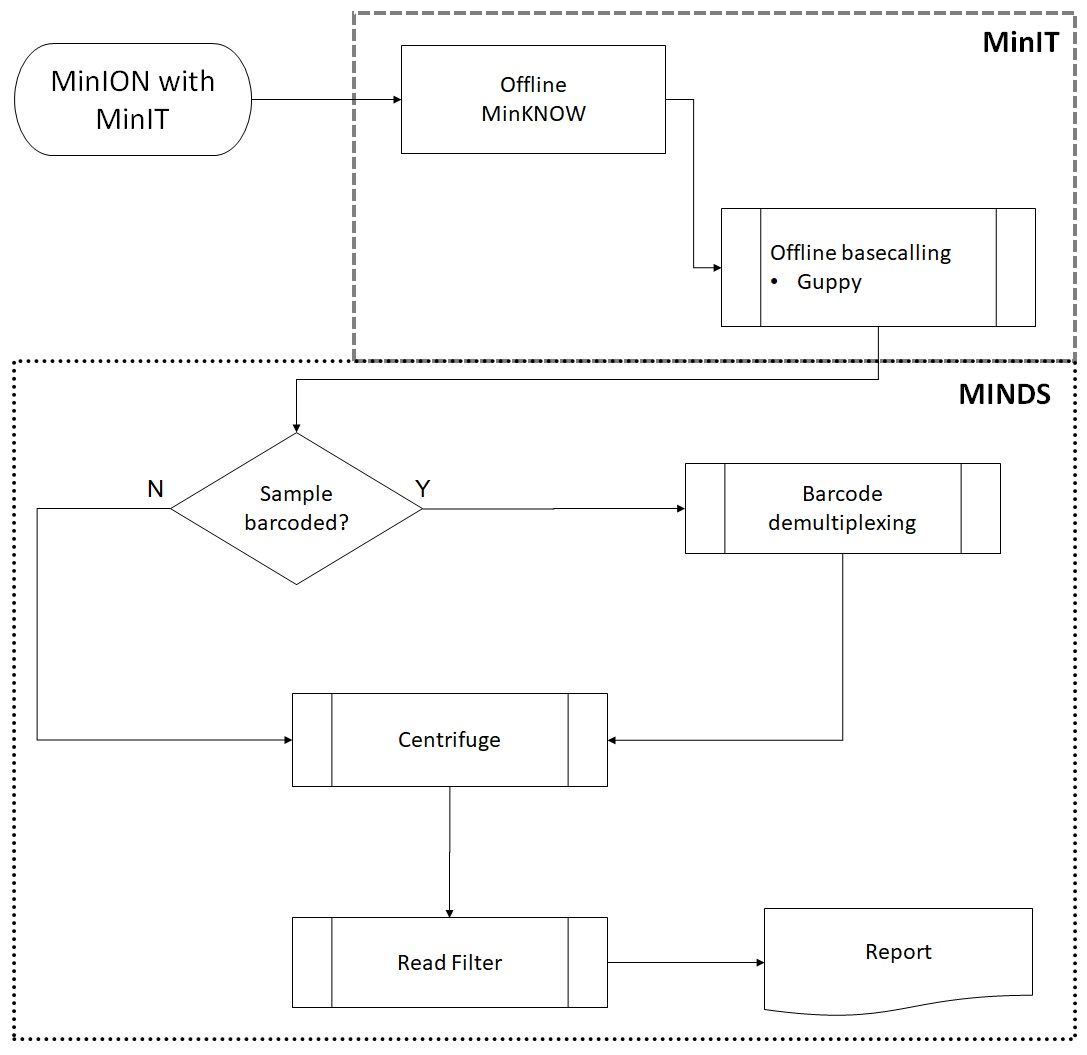
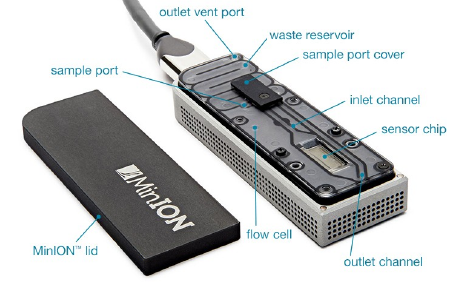
Genes | Free Full-Text | Offline Next Generation Metagenomics Sequence Analysis Using MinION Detection Software (MINDS)
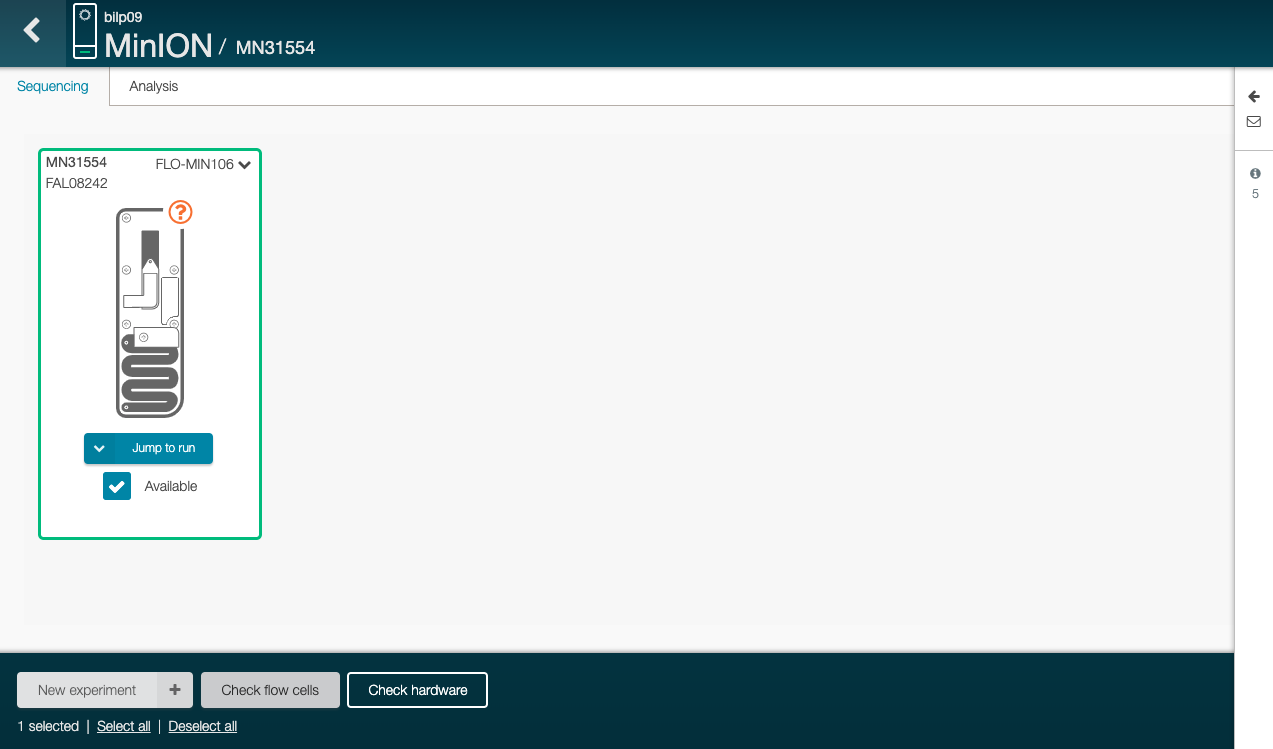
GitHub - envmetagen/nanoenv: Nanopore software - ubuntu image with MinKNOW installed and ready to use with Minion